


Keith Gagnon received his PhD in Biochemistry in 2007 from North Carolina State University. After completing postdoctoral training with David R.Corey at UT Southwestern Medical Center in Dallas, TX, he was jointly appointed to the Department of Chemistry and the Department of Biochemistry and Molecular Biology in the School of Medicine in 2014.
(618) 453-3895
email: kgagnon@siumed.edu
RNA is one of the most fascinating molecules in the cell. Hands down. It seems there is no job that RNA doesn't have a stake in, from transcription to translation, catalysis to structure, information storage to ligand binding -- designed by nature to be versatile and multipurpose. Having so many irons in the fire, however, means that RNA also plays key roles in countless forms of disease.
We are interested in the biology and biochemistry of RNA, especially non-coding RNA, and how its sequence and structure correlate with function. We also want to know what goes wrong when RNA causes or facilitates disease. Our main focus is on non-coding RNA biology in the mammalian cell nucleus. Our main diseases of interest are the growing class of neurological disorders known as repeat expansion diseases. These diseases are caused when naturally occurring repeats in the human genome expand and become pathogenic.
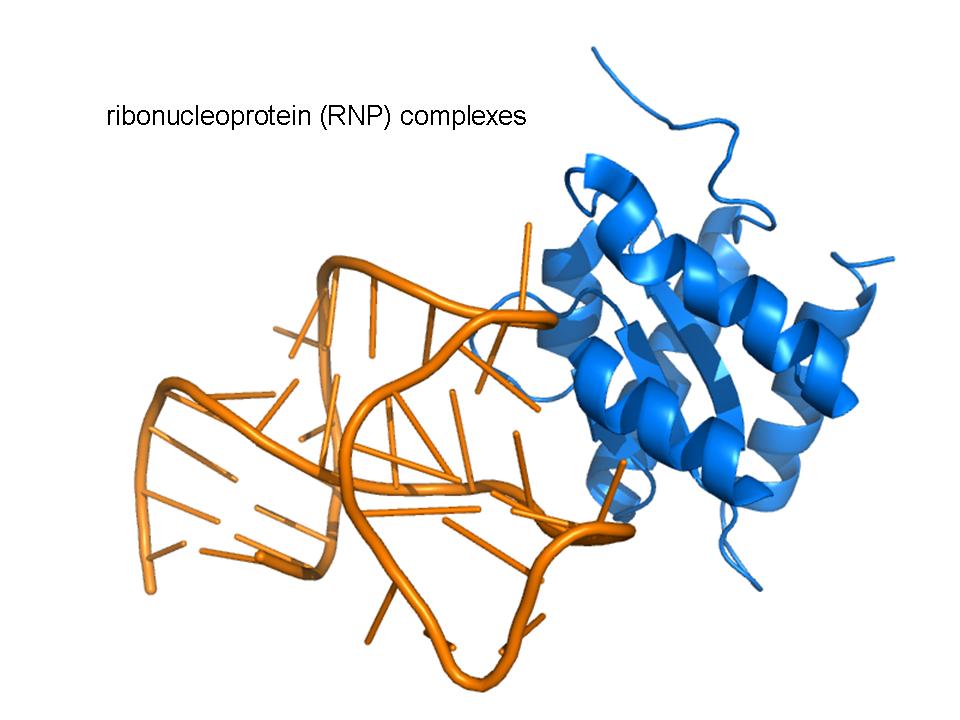
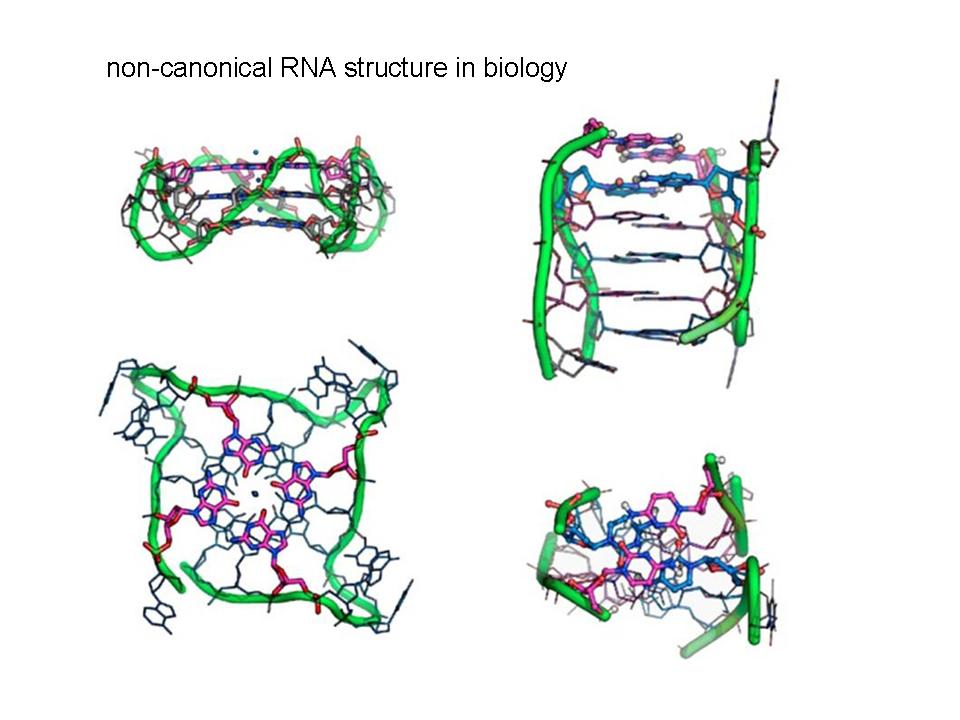
We care about the molecular-level pathology of repeat expansion diseases. Currently, we are focused on a recent newcomer to the family, a repeat expansion disease known as c9FTD/ALS. This disease is a spectrum disorder that causes amyotrophic lateral sclerosis (ALS, or Lou Gehrig's Disease) and frontotemporal dementia (FTD) and may play a role in other neurological disorders. c9FTD/ALS is caused by the expansion of a hexanucleotide repeat in an intron of the C9ORF72 gene that is transcribed into large repeat RNAs. These RNAs form foci in the nucleus of cells and are expected to form complicated non-canonical RNA structures and sequester RNA-binding proteins.
We focus our efforts on three fronts, i) understanding the molecular biology and biochemistry of these disorders, ii) developing tools to discover and detect repeat expansions and biomarkers for disease diagnosis, and iii) identifying points for therapeutic intervention in these diseases.
Projects will use a number of approaches, including in vitro biochemistry, biophysics instrumentation, structural analyses (crystallography), mass spectrometry, RNA sequencing, mammalian tissue culture, classic biochemical purifications, enzyme activity assays, microscopy, animal models (in collaboration), nucleic acid chemistry.
We want to discover and characterize nuclear non-coding RNAs (ncRNAs), including small RNAs and long ncRNAs. The rich history of ncRNA function and diversity suggests that many more discoveries are waiting in the nucleus.
Of particular interest are novel roles for long ncRNAs that may act on the chromatin. A common theme for ncRNA that lncRNAs are expected to follow is serving in the roles of structural scaffolds to assemble higher-order ribonucleoprotein complexes and as guides to direct the activity and function of associated proteins.
Projects will use various techniques, including in vitro biochemistry, biophysics instrumentation, structural analyses (crystallography), mass spectrometry, RNA sequencing, mammalian tissue culture, classic biochemical purifications, enzyme activity assays, mutagenesis studies, nucleic acid chemistry, and microscopy.
Gagnon, K.T., Li, L., Chu, Y., Janowski, B.A., and Corey, D.R. (2014) RNAi Factors Are Present and Active in Human Cell Nuclei. Cell Reports, 6:211-221.
Matsui, M., Zhang, H., Chu, J., Gagnon, K.T., Shaikh, S., Kuchimanchi, S., Manoharan, M., Corey, D.R., and Janowski, B.A. (2013) Promoter RNA Links Transcriptional Regulation of Inflammatory Pathway Genes. Nucl. Acids Res., 41:10086-10109. **"Breakthrough" article; deemed by Editors to be in the top 1% of articles accepted by NAR.**
Dodd, D.W., Gagnon, K.T., and Corey, D.R. (2013) Digital Quantitation of Potential Therapeutic Target RNAs. Nucl. Acid Therap., 23:188-194.
Gagnon, K.T., Biswas, S., Zhang, X., Brown, B.A. II, Wollenzien, P., Mattos, C., and Maxwell, E.S. (2012) Structurally Conserved Nop56/58 N-Terminal Domain Facilitates Efficient Box C/D Ribonucleoprotein-guided Methyltransferase Activity. J. Biol. Chem., 287:19418-19428.
Gagnon, K.T., Watts, J.K., Pendergraff, H.M., Potier, P., Thai, D., Montaillier, C., and Corey, D.R. (2011) Antisense and antigene inhibition of gene expression by cell-permeable oligonucleotide-oligospermine conjugates. J. Am. Chem. Soc., 133:8404-8407.
Biswas, S., Buhrman, G., Gagnon, K.T., Mattos, C., Brown II, B.A., and Maxwell, E.S. (2011) Comparative Analysis of the 15.5kD Box C/D snoRNP Core Protein in the Primitive Eukaryote Giardia lamblia Reveals Unique Structural and Functional Features. Biochemistry, 50:2907-2918.
Hu, J., Gagnon, K.T., Lui, J., Watts, J.K., Syeda-Newaz, J., Bennett, C.F., Swayze, E.E., Randolph, J., Chattopadhyaya, J. and Corey, D.R. (2011) Allele-selective inhibition of ataxin-3 (ATX3) expression by antisense oligomers and duplex RNAs. Biol. Chem., 392:315-325.
Gagnon, K.T., Pendergraff, H.M., Deleavey, G.F., Swayze, E.E., Potier, P., Randolph, J., Roesch, E.B., Chattopadhyaya, J., Damha, M.J., Bennett, C.F., Montaillier, C., Lemaitre, M., and Corey, D.R. (2010) Allele-Selective Inhibition of Mutant Huntingtin Expression with Antisense Oligonucleotides Targeting the Expanded CAG Repeat. Biochemistry, 49:10166-10178..
Yue, X., Schwartz, J. C., Chu, Y., Younger, S. T., Gagnon, K.T., Elbashir, S., Janowski, B. A., and Corey, D. R. (2010) Transcriptional regulation by small RNAs at sequences downstream from 3' gene termini. Nat. Chem. Biol., 6:621-629..
Gagnon, K.T., Zhang, X., Qu, G., Biswas, S., Suryadi, J., Brown II, B.A., and Maxwell, E.S. (2010) Signature Amino Acids Enable the Archaeal L7Ae Box C/D RNP Core Protein to Recognize and Bind the K-loop RNA Motif. RNA, 16:79-90.
Bleichert, F., Gagnon, K.T., Brown II, B.A., Maxwell, E.S., Leschziner, A.E., Unger, V.M., and Baserga, S.J. (2009) A Dimeric Structure for Archaeal Box C/D Small Ribonucleoproteins. Science, 325(5946):1384-1387.
Gagnon, K.T., Ju, S.Y., Goshe, M.B., Maxwell, E.S., and Franzen, S. (2009) A Role for Hydrophobicity in a Diels-Alder Reaction Catalyzed by Pyridyl-Modified RNA. Nucl. Acids Res., 37(9):3074-3082.
Hu, J., Matsui, M., Gagnon, K.T., Schwartz, J.C., Gabillet, S., Arar, K., Wu, J., Bezprozvanny, I., and Corey, D.R. (2009) Allele-Specific Silencing of Mutant Huntingtin and Ataxin-3 Genes by Targeting Expanded CAG Repeats in mRNAs. Nat. Biotechnol., 27(5):478-484.
Gagnon, K.T., Zhang, X., Agris, P.F., and Maxwell, E.S. (2006) Assembly of the Archaeal Box C/D sRNP Can Occur Via Alternative Pathways and Requires Temperature-Facilitated sRNA Remodeling. J. Mol. Biol., 362(5):1025-1042.
![]() Biochemistry and Molecular
Biology Home Page
Biochemistry and Molecular
Biology Home Page